Abstract
Mutations in the MYBPC3 gene, encoding the sarcomere protein myosin-binding protein C, are among the most frequent causes of autosomal dominant familial hypertrophic cardiomyopathy (FHC). We studied the frequency, type, and pathogenetic mechanism of MYBPC3 mutations in an unselected cohort of 81 FHC families, consecutively enrolled at a tertiary referral center. Nine mutations, six of which were novel, were found in 10 (12.3%) of the families using single-strand conformation polymorphism and DNA sequencing. A frameshift mutation in exon 2 clearly suggests that haploinsufficiency is a pathogenetic mechanism in FHC. In addition, splice site mutations in exon 6 and intron 31, a deletion in exon 13, and a nonsense mutation in exon 25, all lead to premature termination codons, most likely causing loss of function and haploinsufficiency. Furthermore, there were two missense mutations (D228N and A833 T) and one in-frame deletion (ΔLys813). A considerable intrafamilial variation in phenotypic expression of MYBPC3-based FHC was noted, and we suggest that mutations influencing stability of mRNA could play a role in the variable penetrance and expressivity of the disease, perhaps via partial haploinsuffciency.
Similar content being viewed by others
Introduction
Familial hypertrophic cardiomyopathy (FHC) is an autosomal dominant cardiac disorder characterized by unexplained myocardial hypertrophy, predominantly of the interventricular septum.1, 2 The clinical phenotype varies from asymptomatic over limited symptoms to severe heart failure or the occurrence of serious arrhythmias or sudden cardiac death.3
FHC has been associated with more than 200 mutations in 10 known genes encoding cardiac sarcomere proteins.4 Furthermore, mutations in other genes (eg mitochondrial DNA and PRKAG2) have been associated with a cardiac phenotype resembling hypertrophic cardiomyopathy.5, 6 Mutations in MYH7, encoding β-myosin heavy chain, and in MYBPC3, encoding myosin binding protein C, are the most prevalent and probably account for 20–40% of FHC cases.4, 7
We studied the occurrence of MYBPC3 mutations in a cohort of 81 consecutive patients in a nationwide Danish FHC study. We found nine mutations, six of which were novel, in 10 families. In three families, mutations in both MYBPC3 and MYH7 were found. Several of the mutations undoubtedly result in total or partial haploinsufficiency.
Materials and methods
Patients
In all, 81 unrelated FHC families were remitted consecutively from The National University Hospital, Rigshospitalet, Copenhagen, a tertiary referral center for FHC. DNA samples from 100 blood donor samples were used as controls.
Clinical evaluation
Clinical examinations were performed as previously described.8 All probands fulfilled conventional diagnostic criteria for hypertrophic cardiomyopathy.9
Mutation analyses
Mutation analyses were performed by single-strand conformation polymorphism/heteroduplex analysis (SSCP/HA)10 or capillary array single-strand conformation polymorphism (CAE-SSCP)11, 12 of amplicons covering all 34 coding exons of MYBPC3 using intronic primers from13 or defined from the gene sequence (GenBank Accession No. U91629). The sequence of primers and PCR conditions are available upon request. Genetic variants with frequencies above 1% in controls were considered polymorphisms. Seven other FHC-associated genes (ACTC, MYH7, MYL2, MYL3, TNNI3, TNNT2, and TPM1)11, 14, 15 were screened for mutations in the genotyped probands.
Messenger RNA transcript analysis
Ectopic mRNA expression of MYBPC3 in peripheral blood lymphocytes was studied.16, 17, 18 RNA was extracted from fresh blood using an RNA purification kit (Qiagen, Germany). RT-PCR was performed using MMLV Reverse Transcriptase (Stratagene, LA Jolla, CA, USA) and exon specific primers. In case of amplification with the 5′-proximal primer set, MYBPC1EF (5′-CGACCAGGGATCTTACGCATG-3′) and MYBPC12ER (5′-GGGTGCCTGCCGTAGGATCTC-3′) a second, nested PCR amplification was applied on the primary amplicon using primers MYBPC294F (5′-CGACCAGGGATCTTACGCAGT-3′) and MYBPC897R (5′-CCTCA TGGCTATCACTGATCCG-3′). For amplification of the 3′-proximal region, primers MYBPC29EF (5′-CGCTCGCC GCGTGCATTCAG-3′) and MYBPC33ER (5′-CCGTCAAAG GGGCAGGGCTTTC-3′) were applied. RT-PCR products were cloned into pCR-Script (SK) (Stratagene) and sequenced.
Results
Mutation screening
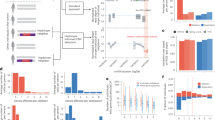
Nine mutations were detected in 10 families with one mutation present in two families (Table 1). Three of the mutations (g5256G>A (Glu258Lys; Figure 1), g16153G>A (Ala833Thr), and g15919insG) have previously been characterized.13, 19 The six novel mutations consisted of a missense mutation (g5166G>A (Asp228Asn)) and a nonsense mutation (g16182C>G; Tyr842Ter), three exonic deletions (g2432delT, g10080-96delCTGACCGTGGAACTGG, and g16086delAAG (DelLys813) and one intronic deletion (g20931-37delCCTGTCA(−8-(−)14)).
Detection of g5256G>A (Glu258Lys) mutation and consequent exon skipping. (a) Capillary array SSCP, where the upper two panels represent the wild-type chromatograms (forward (upper) and reverse (lower), respectively); the two lower panels represent the mutant chromatograms (forward (upper) and reverse (lower), respectively); (b) DNA sequence of the wild-type control (upper panel) and the heterozygote (G>A substitution) (lower panel); (c) DNA sequence of RT-PCR clone containing the splice site mutation; upper panel illustrates sequence where exon 6 has been skipped and lower panel the normal cDNA sequence.
The small size of most of the genotyped families makes it difficult to establish a causal relationship between the occurrence of a specific mutation and FHC. MYBPC3 and MYH7 mutations were found in the families E, G and XI, where the latter two carried very rare variants in MYBPC3 of unknown significance besides disease-associated mutations in MYH7.
Messenger RNA studies
In order to determine the consequences of mutations, we performed analysis of ectopically expressed MYBPC3 mRNA in the absence of cardiac tissue. Nested RT-PCR performed on DNA from a g2432ΔT mutation carrier using exon-specific primers MYBPC1EF/MYBPC12ER followed by nested PCR with primers MYBPC294F/MYBPC897R) resulted in one band on a gel, migrating as an approximately 500 bp DNA fragment. Cloning and sequencing revealed that both alleles were present.
Nested RT-PCR analysis in a g5256 mutation carrier, from family ZI, using the primer pair MYBP294F and MYBPC897R for reamplification, yielded two dominant PCR products corresponding to the wild-type allele and an allele with exon 6 skipping (Figure 1c). Exon skipping resulted in a frameshift and a premature termination codon, and, most likely as a consequence in haploinsufficiency.
The deletion in intron 31 (g20931-37delCCTGTCA (−8(−)14)) was shown by RT-PCR using MYBPC29F and MYBPC33R primers to result in both a normally spliced transcript and a transcript with exon 32 skipped.
The remaining splice-site mutations with a possible splice variation, IVS8-20C>A and IVS16-6G>A, and the insertion/deletion mutations could not be analyzed as fresh blood or cardiac tissue from the mutation carriers could not be obtained.
Clinical evaluation of patients with MYBPC3 mutations
Among the 44 carriers with mutations solely in MYBPC3, there were 25 carriers (57%) that had a maximum LVD⩾13 mm and 14 carriers (32%) with HCM symptoms or prior myectomy. In three families, ZF, ZH, and ZP, no major cardiac events had occurred, and in the other families, one to two members had experienced a cardiac event. Three of the probands with a MYBPC3 mutation (C, ZL, and ZV) had a debut age under 18 years with severe hypertrophy. A detailed clinical description of patients with digenic inheritance is shown in Table 2. No specific phenotype could be attributed to these patients.
Discussion
The prevalence of MYBPC3 mutations in Denmark, 10/81 (12.3%) is similar to the frequency of 13.6% (15/110) recently reported in a study of Caucasian patients20 and 10.8% (14/130) in a Japanese study13 but much lower than recent reports where 24% of Finnish HCM cases carried MYBPC3 mutations21 and in a French study with a prevalence of 42% MYBPC3 mutations among 124 genotyped patients.4
In agreement with other studies, we find that late onset is characteristic, but in some families, children may be affected.13, 20 The relatively low proportion of mutations leading to single amino-acid substitutions is similar to the findings in the Caucasian study,20 contrary to the findings in a Japanese and a recent French study.4, 13 The g15919insG mutation, previously described by Niimura et al,13 was found in two families (ZF and ZP) and could be a founder mutation, analogous to the Turkish, Finnish and South African founder mutations.20, 22 Haplotype analysis using STRs in the flanking regions as well as SNP analysis in the MYBPC3 shows that the two probands are not closely related.23
The finding of three families with mutations in both MYBPC3 and MYH7 is surprising, but the phenomenon of digenic inheritance in these genes has previously been described in recent French and European studies.4, 24 These findings should lead to a re-evaluation of all patients diagnosed with HCM based on MYBPC3 mutations, since they may carry the true disease-causing mutations on other genes.
Haploinsufficiency
Six out of the 10 mutations found in this study resulted in generation of a premature termination codon (Table 1). For at least three of these (g2432ΔT, g5256G>A, and g10080-96delCT…GG), and possibly also the remaining three, the translation product will be so short as to prevent effectively its incorporation in the sarcomere; thus in these three cases, there will be a quantitative rather than a qualitative deficiency of the sarcomere – haploinsufficiency – as mRNAs containing premature termination codons are often rapidly degraded through the so-called nonsense mediated mRNA decay.25 However, the splice variants that were observed may be artifacts due to ectopic expression in leukocytes, and final proof of aberrant splicing for mRNA or protein studies may only be documented in cardiac tissue, which has not been available in the present cases.
It is interesting that the most seriously affected families, ZV, ZI, and ZL are the ones where haploinsufficiency as a pathogenic mechanism is mostly likely. However, only large studies comparing clinical phenotypes with mutations and polymorphisms, in large families, and corresponding studies of transcription and translation, can provide the certain knowledge we need to have in order to use the genetic workup clinically. Presently, as our study documents, mutations in MYBPC3 should be studied carefully before being assigned as disease causing.
References
Maron BJ, Bonow RO, Cannon RO et al: Hypertrophic cardiomyopathy. Interrelations of clinical manifestations, pathophysiology, and therapy (2). N Engl J Med 1987; 316: 844–852.
Wigle ED, Rakowski H, Kimball BP et al: Hypertrophic cardiomyopathy. Clinical spectrum and treatment. Circulation 1995; 92: 1680–1692.
Spirito P, Seidman CE, McKenna WJ et al: The management of hypertrophic cardiomyopathy. N Engl J Med 1997; 336: 775–785.
Richard P, Charron P, Carrier L et al: Hypertrophic cardiomyopathy: distribution of disease genes, spectrum of mutations, and implications for a molecular diagnosis strategy. Circulation 2003; 107: 2227–2232.
Merante F, Myint T, Tein I et al: An additional mitochondrial tRNA(Ile) point mutation (A-to-G at nucleotide 4295) causing hypertrophic cardiomyopathy. Hum Mutat 1996; 8: 216–222.
Arad M, Benson DW, Perez-Atayde AR et al: Constitutively active AMP kinase mutations cause glycogen storage disease mimicking hypertrophic cardiomyopathy. J Clin Invest 2002; 109: 357–362.
Seidman JG, Seidman C : Transcription factor haploinsufficiency: when half a loaf is not enough. J Clin Invest 2002; 109: 451–455.
Havndrup O, Bundgaard H, Skytt AP et al: Outcome of clinical versus genetic family screening in hypertrophic cardiomyopathy with focus on cardiac beta-myosin gene mutations. Cardiovasc Res 2003; 57: 347–357.
Maron BJ, Gottdiener JS, Epstein SE : Patterns and significance of distribution of left ventricular hypertrophy in hypertrophic cardiomyopathy. A wide angle, two dimensional echocardiographic study of 125 patients. Am J Cardiol 1981; 48: 418–428.
Bundgaard H, Havndrup O, Andersen PS et al: Familial hypertrophic cardiomyopathy associated with a novel missense mutation affecting the ATP-binding region of the cardiac beta-myosin heavy chain. J Mol Cell Cardiol 1999; 31: 745–750.
Larsen LA, Christiansen M, Vuust J et al: High-throughput single-strand conformation polymorphism analysis by automated capillary electrophoresis: robust multiplex analysis and pattern-based identification of allelic variants. Hum Mutat 1999; 13: 318–327.
Andersen PS, Jespersgaard C, Vuust J et al: High-throughput single strand conformation polymorphism mutation detection by automated capillary array electrophoresis: validation of the method. Hum Mutat 2003; 21: 116–122.
Niimura H, Bachinski LL, Sangwatanaroj S et al: Mutations in the gene for cardiac myosin-binding protein C and late- onset familial hypertrophic cardiomyopathy. N Engl J Med 1998; 338: 1248–1257.
Andersen PS, Havndrup O, Bundgaard H et al: Adult-onset familial hypertrophic cardiomyopathy caused by a novel mutation, R694C, in the MYH7 gene. Clin Genet 1999; 56: 244–246.
Andersen PS, Havndrup O, Bundgaard H et al: Myosin light chain mutations in familial hypertrophic cardiomyopathy: phenotypic presentation and frequency in Danish and South African populations. J Med Genet 2001; 38: E43.
Sarkar G, Sommer SS : Access to a messenger RNA sequence or its protein product is not limited by tissue or species specificity. Science 1989; 244: 331–334.
Rosenzweig A, Watkins H, Hwang DS et al: Preclinical diagnosis of familial hypertrophic cardiomyopathy by genetic analysis of blood lymphocytes. N Engl J Med 1991; 325: 1753–1760.
Watkins H, Rosenzweig A, Hwang DS et al: Characteristics and prognostic implications of myosin missense mutations in familial hypertrophic cardiomyopathy. N Engl J Med 1992; 326: 1108–1114.
Morner S, Richard P, Kazzam E et al: Identification of the genotypes causing hypertrophic cardiomyopathy in northern Sweden. J Mol Cell Cardiol 2003; 35: 841–849.
Erdmann J, Raible J, Maki-Abadi J et al: Spectrum of clinical phenotypes and gene variants in cardiac myosin-binding protein C mutation carriers with hypertrophic cardiomyopathy. J Am Coll Cardiol 2001; 38: 322–330.
Jaaskelainen P, Kuusisto J, Miettinen R et al: Mutations in the cardiac myosin-binding protein C gene are the predominant cause of familial hypertrophic cardiomyopathy in eastern Finland. J Mol Med 2002; 80: 412–422.
Moolman-Smook JC, De Lange WJ, Bruwer EC et al: The origins of hypertrophic cardiomyopathy-causing mutations in two South African subpopulations: a unique profile of both independent and founder events. Am J Hum Genet 1999; 65: 1308–1320.
Mogensen J, Andersen PS, Steffensen U, Christiansen M, Gregersen N, Børglum AD : Development and application of linkage analysis in genetic diagnosis of familial hypertrophic cardiomyopathy. J Med Genet 2001; 38: 193–197.
Richard P, Isnard R, Carrier L et al: Double heterozygosity for mutations in the beta-myosin heavy chain and in the cardiac myosin binding protein C genes in a family with hypertrophic cardiomyopathy. J Med Genet 1999; 36: 542–545.
Byers PH : Killing the messenger: new insights into nonsense-mediated mRNA decay. J Clin Invest 2002; 109: 3–6.
Acknowledgements
We thank Anders Børglum, MD, for his discussions in this study, and we gratefully acknowledge the technical assistance of Kirsten Lindboe, Mads Dahm Johansen, and Jette Severinsen. The study has been supported by grants from The Danish Heart Foundation, Director Emil C Hertz and wife Inger Hertz's foundation, Danish Medical Association Research Foundation, H:S – Copenhagen Hospital Corporation, “The John and Birte Meyer Foundation” and The Danish Medical Research Council; Copenhagen, Denmark.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Andersen, P., Havndrup, O., Bundgaard, H. et al. Genetic and phenotypic characterization of mutations in myosin-binding protein C (MYBPC3) in 81 families with familial hypertrophic cardiomyopathy: total or partial haploinsufficiency. Eur J Hum Genet 12, 673–677 (2004). https://doi.org/10.1038/sj.ejhg.5201190
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201190
Keywords
This article is cited by
-
Allelic imbalance and haploinsufficiency in MYBPC3-linked hypertrophic cardiomyopathy
Pflügers Archiv - European Journal of Physiology (2019)
-
Successful knock-in of Hypertrophic Cardiomyopathy-mutation R723G into the MYH7 gene mimics HCM pathology in pigs
Scientific Reports (2018)
-
Clinical outcomes associated with sarcomere mutations in hypertrophic cardiomyopathy: a meta-analysis on 7675 individuals
Clinical Research in Cardiology (2018)
-
Evaluation of the Mayo Clinic Phenotype-Based Genotype Predictor Score in Patients with Clinically Diagnosed Hypertrophic Cardiomyopathy
Journal of Cardiovascular Translational Research (2016)
-
Screening Mutations of MYBPC3 in 114 Unrelated Patients with Hypertrophic Cardiomyopathy by Targeted Capture and Next-generation Sequencing
Scientific Reports (2015)