Abstract
Although de novo missense mutations have been predicted to account for more cases of autism than gene-truncating mutations, most research has focused on the latter. We identified the properties of de novo missense mutations in patients with neurodevelopmental disorders (NDDs) and highlight 35 genes with excess missense mutations. Additionally, 40 amino acid sites were recurrently mutated in 36 genes, and targeted sequencing of 20 sites in 17,688 patients with NDD identified 21 new patients with identical missense mutations. One recurrent site substitution (p.A636T) occurs in a glutamate receptor subunit, GRIA1. This same amino acid substitution in the homologous but distinct mouse glutamate receptor subunit Grid2 is associated with Lurcher ataxia. Phenotypic follow-up in five individuals with GRIA1 mutations shows evidence of specific learning disabilities and autism. Overall, we find significant clustering of de novo mutations in 200 genes, highlighting specific functional domains and synaptic candidate genes important in NDD pathology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Ronemus, M., Iossifov, I., Levy, D. & Wigler, M. The role of de novo mutations in the genetics of autism spectrum disorders. Nat. Rev. Genet. 15, 133–141 (2014).
Bernier, R. et al. Disruptive CHD8 mutations define a subtype of autism early in development. Cell 158, 263–276 (2014).
Stessman, H.A., Bernier, R. & Eichler, E.E. A genotype-first approach to defining the subtypes of a complex disease. Cell 156, 872–877 (2014).
Sanders, S.J. et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 87, 1215–1233 (2015).
Packer, A. Neocortical neurogenesis and the etiology of autism spectrum disorder. Neurosci. Biobehav. Rev. 64, 185–195 (2016).
Turner, T.N. et al. Proteins linked to autosomal dominant and autosomal recessive disorders harbor characteristic rare missense mutation distribution patterns. Hum. Mol. Genet. 24, 5995–6002 (2015).
van Bon, B.W.M. et al. Disruptive de novo mutations of DYRK1A lead to a syndromic form of autism and ID. Mol. Psychiatry 21, 126–132 (2016).
Helsmoortel, C. et al. A SWI/SNF-related autism syndrome caused by de novo mutations in ADNP. Nat. Genet. 46, 380–384 (2014).
Buxbaum, J.D. DSM-5 and psychiatric genetics — round hole, meet square peg. Biol. Psychiatry 77, 766–768 (2015).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Turner, T.N. et al. Genome sequencing of autism-affected families reveals disruption of putative noncoding regulatory DNA. Am. J. Hum. Genet. 98, 58–74 (2016).
O'Roak, B.J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012).
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
McRae, J.F. et al. Prevalence and architecture of de novo mutations in developmental disorders. Nature http://dx.doi.org/10.1038/nature21062 (2017).
Stessman, H.A.F., Turner, T.N. & Eichler, E.E. Molecular subtyping and improved treatment of neurodevelopmental disease. Genome Med. 8, 22 (2016).
Schuurs-Hoeijmakers, J.H.M. et al. Recurrent de novo mutations in PACS1 cause defective cranial-neural-crest migration and define a recognizable intellectual-disability syndrome. Am. J. Hum. Genet. 91, 1122–1127 (2012).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Lynch, M. Rate, molecular spectrum, and consequences of human mutation. Proc. Natl. Acad. Sci. USA 107, 961–968 (2010).
Maheshwari, M. et al. PTPN11 mutations in Noonan syndrome type I: detection of recurrent mutations in exons 3 and 13. Hum. Mutat. 20, 298–304 (2002).
Myhre, S.A., Ruvalcaba, R.H.A. & Graham, C.B. A new growth deficiency syndrome. Clin. Genet. 20, 1–5 (1981).
Le Goff, C. et al. Mutations at a single codon in Mad homology 2 domain of SMAD4 cause Myhre syndrome. Nat. Genet. 44, 85–88 (2011).
Landrum, M.J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D1, D862–D868 (2016).
de Ligt, J. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012).
Yuan, H., Erreger, K., Dravid, S.M. & Traynelis, S.F. Conserved structural and functional control of N-methyl-d-aspartate receptor gating by transmembrane domain M3. J. Biol. Chem. 280, 29708–29716 (2005).
Zuo, J. et al. Neurodegeneration in Lurcher mice caused by mutation in δ2 glutamate receptor gene. Nature 388, 769–773 (1997).
Coutelier, M. et al. GRID2 mutations span from congenital to mild adult-onset cerebellar ataxia. Neurology 84, 1751–1759 (2015).
Kohda, K., Wang, Y. & Yuzaki, M. Mutation of a glutamate receptor motif reveals its role in gating and δ2 receptor channel properties. Nat. Neurosci. 3, 315–322 (2000).
Taverna, F. et al. The Lurcher mutation of an α-amino-3-hydroxy-5-methyl- 4-isoxazolepropionic acid receptor subunit enhances potency of glutamate and converts an antagonist to an agonist. J. Biol. Chem. 275, 8475–8479 (2000).
Klein, R.M. & Howe, J.R. Effects of the lurcher mutation on GluR1 desensitization and activation kinetics. J. Neurosci. 24, 4941–4951 (2004).
Kessels, H.W. & Malinow, R. Synaptic AMPA receptor plasticity and behavior. Neuron 61, 340–350 (2009).
Tavassoli, T. et al. De novo SCN2A splice site mutation in a boy with autism spectrum disorder. BMC Med. Genet. 15, 35 (2014).
Allen, A.S. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Niknafs, N. et al. MuPIT interactive: webserver for mapping variant positions to annotated, interactive 3D structures. Hum. Genet. 132, 1235–1243 (2013).
Banerjee, S., Riordan, M. & Bhat, M.A. Genetic aspects of autism spectrum disorders: insights from animal models. Front. Cell. Neurosci. 8, 58 (2014).
Durand, C.M. et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat. Genet. 39, 25–27 (2007).
Hamdan, F.F. et al. De novo SYNGAP1 mutations in nonsyndromic intellectual disability and autism. Biol. Psychiatry 69, 898–901 (2011).
Peñagarikano, O. et al. Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell 147, 235–246 (2011).
Varoqueaux, F. et al. Neuroligins determine synapse maturation and function. Neuron 51, 741–754 (2006).
Zanjani, H.S. et al. Death and survival of heterozygous Lurcher Purkinje cells in vitro. Dev. Neurobiol. 69, 505–517 (2009).
Andrásfalvy, B.K., Smith, M.A., Borchardt, T., Sprengel, R. & Magee, J.C. Impaired regulation of synaptic strength in hippocampal neurons from GluR1-deficient mice. J. Physiol. (Lond.) 552, 35–45 (2003).
Wiedholz, L.M. et al. Mice lacking the AMPA GluR1 receptor exhibit striatal hyperdopaminergia and 'schizophrenia-related' behaviors. Mol. Psychiatry 13, 631–640 (2008).
Barcia, G. et al. De novo gain-of-function KCNT1 channel mutations cause malignant migrating partial seizures of infancy. Nat. Genet. 44, 1255–1259 (2012).
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
Dimassi, S. et al. Whole-exome sequencing improves the diagnosis yield in sporadic infantile spasm syndrome. Clin. Genet. 89, 198–204 (2016).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Gulsuner, S. et al. Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network. Cell 154, 518–529 (2013).
Hashimoto, R. et al. Whole-exome sequencing and neurite outgrowth analysis in autism spectrum disorder. J. Hum. Genet. 61, 199–206 (2016).
Helbig, K.L. et al. Diagnostic exome sequencing provides a molecular diagnosis for a significant proportion of patients with epilepsy. Genet. Med. 18, 898–905 (2016).
Homsy, J. et al. De novo mutations in congenital heart disease with neurodevelopmental and other congenital anomalies. Science 350, 1262–1266 (2015).
Jiang, Y.H. et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am. J. Hum. Genet. 93, 249–263 (2013).
Kranz, T.M. et al. De novo mutations from sporadic schizophrenia cases highlight important signaling genes in an independent sample. Schizophr. Res. 166, 119–124 (2015).
Krumm, N. et al. Excess of rare, inherited truncating mutations in autism. Nat. Genet. 47, 582–588 (2015).
Lee, H., Lin, M.C., Kornblum, H.I., Papazian, D.M. & Nelson, S.F. Exome sequencing identifies de novo gain of function missense mutation in KCND2 in identical twins with autism and seizures that slows potassium channel inactivation. Hum. Mol. Genet. 23, 3481–3489 (2014).
Lelieveld, S.H. et al. Meta-analysis of 2,104 trios provides support for 10 new genes for intellectual disability. Nat. Neurosci. 19, 1194–1196 (2016).
McCarthy, S.E. et al. De novo mutations in schizophrenia implicate chromatin remodeling and support a genetic overlap with autism and intellectual disability. Mol. Psychiatry 19, 652–658 (2014).
Michaelson, J.J. et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell 151, 1431–1442 (2012).
O'Roak, B.J. et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat. Commun. 5, 5595 (2014).
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012).
Veeramah, K.R. et al. De novo pathogenic SCN8A mutation identified by whole-genome sequencing of a family quartet affected by infantile epileptic encephalopathy and SUDEP. Am. J. Hum. Genet. 90, 502–510 (2012).
Veeramah, K.R. et al. Exome sequencing reveals new causal mutations in children with epileptic encephalopathies. Epilepsia 54, 1270–1281 (2013).
Yuen, R.K.C. et al. Whole-genome sequencing of quartet families with autism spectrum disorder. Nat. Med. 21, 185–191 (2015).
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013).
Turner, T.N. et al. Denovo-db: a compendium of human de novo variants. Nucleic Acids Res. 45, D1, D804–D811 (2017).
Genome of the Netherlands Consortium. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat. Genet. 46, 818–825 (2014).
Constantino, J. Social Responsiveness Scale (SRS-2). West. Psychol. Serv. (2012).
Ng, S.B. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Ezkurdia, I. et al. Multiple evidence strands suggest that there may be as few as 19,000 human protein-coding genes. Hum. Mol. Genet. 23, 5866–5878 (2014).
Pirooznia, M. et al. SynaptomeDB: an ontology-based knowledgebase for synaptic genes. Bioinformatics 28, 897–899 (2012).
Subtil-Rodríguez, A. et al. The chromatin remodeller CHD8 is required for E2F-dependent transcription activation of S-phase genes. Nucleic Acids Res. 42, 2185–2196 (2014).
Darnell, J.C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Hiatt, J.B., Pritchard, C.C., Salipante, S.J., O'Roak, B.J. & Shendure, J. Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation. Genome Res. 23, 843–854 (2013).
Boyle, E.A., O'Roak, B.J., Martin, B.K., Kumar, A. & Shendure, J. MIPgen: optimized modeling and design of molecular inversion probes for targeted resequencing. Bioinformatics 30, 2670–2672 (2014).
Hoischen, A. et al. De novo mutations of SETBP1 cause Schinzel-Giedion syndrome. Nat. Genet. 42, 483–485 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Girirajan, S. et al. Refinement and discovery of new hotspots of copy-number variation associated with autism spectrum disorder. Am. J. Hum. Genet. 92, 221–237 (2013).
Coe, B.P. et al. Refining analyses of copy number variation identifies specific genes associated with developmental delay. Nat. Genet. 46, 1063–1071 (2014).
Moreno-Ramos, O.A., Olivares, A.M., Haider, N.B., de Autismo, L.C. & Lattig, M.C. Whole-exome sequencing in a south American cohort links ALDH1A3, FOXN1 and retinoic acid regulation pathways to autism spectrum disorders. PLoS One 10, e0135927 (2015).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Douville, C. et al. CRAVAT: cancer-related analysis of variants toolkit. Bioinformatics 29, 647–648 (2013).
Acknowledgements
We thank the individuals and their families for participation in this study. We are grateful to all of the families at the participating Simons Simplex Collection (SSC) sites, as well as the principal investigators (A. Beaudet, R. Bernier, J. Constantino, E. Cook, E. Fombonne, D. Geschwind, R. Goin-Kochel, E. Hanson, D. Grice, A. Klin, D. Ledbetter, C. Lord, C. Martin, D. Martin, R. Maxim, J. Miles, O. Ousley, K. Pelphrey, B. Peterson, J. Piggot, C. Saulnier, M. State, W. Stone, J. Sutcliffe, C. Walsh, Z. Warren, E. Wijsman). We appreciate obtaining access to phenotypic data on SFARI Base. Approved researchers can obtain the SSC population data set described in this study (http://sfari.org/resources/simons-simplex-collection) by applying at https://base.sfari.org/. We gratefully acknowledge the resources provided by the Autism Genetic Resource Exchange (AGRE) Consortium and the participating AGRE families. AGRE is a program of Autism Speaks and is supported, in part, by grant 1U24MH081810 from the National Institute of Mental Health to principal investigator C.M. Lajonchere. We thank J. Gerdts, S. Trinh and B. McKenna for their contributions and T. Brown for assistance in editing this manuscript. This research was supported, in part, by the following: Simons Foundation Autism Research Initiative (SFARI 303241) to E.E.E., National Institutes of Health (R01MH101221 to E.E.E., R01MH100047 to R.A.B., R01MH104450 to L.S.Z., RO1MH105527 and R01DC014489 to J.J.M.), an NHGRI Interdisciplinary Training in Genome Science Grant (T32HG00035) to H.A.F.S. and T.N.T., postdoctoral fellowship grant from the Autism Science Foundation (16-008) to T.N.T., Australian NHMRC grants 1091593 and 1041920 and Channel 7 Children's Research Foundation support to J.G., the National Basic Research Program of China (2012CB517900) and the National Natural Science Foundation of China (81330027, 81525007 and 31671114) to K.X. and H.G., the China Scholarship Council (201406370028) and the Fundamental Research Funds for the Central Universities (2012zzts110) to T.W., grants from the Jack Brockhoff Foundation and Perpetual Trustees, the Victorian State Government Operational Infrastructure Support and Australian Government NHMRC IRIISS, the Swedish Brain Foundation, the Swedish Research Council, the Stockholm County Council, grants (KL2TR00099 and 1KL2TR001444) from the University of California, San Diego Clinical and Translational Research Institute to T.P., the Research Fund – Flanders (FWO) to R.F.K. and G.V., Czech Republic Grant 00064203 and Norway Grant NF-CZ11-PDP-3-003-2014 to Z.S., and the Italian Ministry of Health and '5 per mille' funding to C.R. H.P. is a Senior Clinical Investigator of The Research Foundation–Flanders (FWO). E.E.E. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
E.E.E., L.S.Z., M.R.G., G.H., T.N.T., B.P.C., H.A.F.S. and K.X. designed the study; M.R.G., G.H., T.N.T., B.P.C., T.W. and K.H. performed the experiments; B.P.C. and T.N.T. helped with MIP design and data analysis; M.K., M.N., M.S., J.G., C.B., E.M.T., G.V., R.F.K., T.P., S.N., H.P., C.R., R.A.B., K.X. and H.H. tested inheritance and provided clinical follow-up on select patients; other authors participated in the sample collection and DNA extraction and/or preparation. M.R.G., E.E.E., L.S.Z., G.H., B.P.C. and T.N.T. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
E.E.E. is on the scientific advisory board of DNAnexus, Inc.
Integrated supplementary information
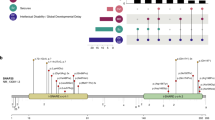
Supplementary Figure 1 Missense mutation clustering in PTPN11.
a) Linear representation of the protein (NP_002825.3) with known functional domains annotated. The top two lines show mutations in controls from the 1000 Genomes Project (1KG)79 and the Exome Aggregation Consortium (ExAC)18. The third line shows de novo missense mutations in cases in denovo-db64 v.1.2. The case mutations fall in three small clusters. b) 3D representation of the PTPN11 protein shows that the three clusters of mutations that are far apart in the linear plot are in close proximity to each other and the ligand binding site after folding34.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1, Supplementary Tables 1 and 11, and Supplementary Clinical Case Reports (PDF 499 kb)
Supplementary Table 2
Genes with recurrent de novo missense mutations in individuals with neurodevelopmental disorders (XLSX 110 kb)
Supplementary Table 3
Genes with recurrent de novo missense mutations in unaffected controls (XLSX 27 kb)
Supplementary Table 4
Recurrent de novo missense mutations at specific amino acid sites (XLSX 29 kb)
Supplementary Table 5
Cohorts sequenced with smMIPs (XLSX 20 kb)
Supplementary Table 6
Regions and cohorts targeted with smMIPs (XLSX 25 kb)
Supplementary Table 7
Rare missense variants with CADD >20 identified with targeted sequencing (XLSX 58 kb)
Supplementary Table 8
All new de novo missense mutations in NDD cases (XLSX 23 kb)
Supplementary Table 9
Phenotypes in cases with GRIA1 p.A636T mutation (XLSX 18 kb)
Supplementary Table 10
All genes with significant (P < 0.05) clustering of de novo missense mutations by CLUMP (XLSX 43 kb)
Rights and permissions
About this article
Cite this article
Geisheker, M., Heymann, G., Wang, T. et al. Hotspots of missense mutation identify neurodevelopmental disorder genes and functional domains. Nat Neurosci 20, 1043–1051 (2017). https://doi.org/10.1038/nn.4589
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4589
This article is cited by
-
A multi-ancestry genetic study of pain intensity in 598,339 veterans
Nature Medicine (2024)
-
Next-generation sequencing and bioinformatics in rare movement disorders
Nature Reviews Neurology (2024)
-
Establishment and characterization of ZJUCHi003: an induced pluripotent stem cell line from a patient with Temple–Baraitser/Zimmermann–Laband syndrome carrying KCNH1 c.1070G > A (p.R357Q) variant
Human Cell (2024)
-
Exploring key genes and pathways associated with sex differences in autism spectrum disorder: integrated bioinformatic analysis
Mammalian Genome (2024)
-
Preliminary study on toxicological mechanism of golden cuttlefish (Sepia esculenta) larvae exposed to cd
BMC Genomics (2023)